Fragile X protein found to regulate key autism candidates
The protein missing in people with fragile X syndrome regulates the activity of more than 800 other proteins, including some key players in autism, according to a study published 22 July in Cell. Many of these autism-associated proteins cluster on either side of the synapse, the junction between neurons.
FMRP, the protein missing in fragile X syndrome, regulates the activity of more than 800 other proteins, including some key players in autism, according to a study published last week in Cell1. Many of these autism-associated proteins cluster on either side of the synapse, the junction between neurons.
Fragile X syndrome is an inherited form of mental retardation caused by a full or partial loss of function of the FMR1 gene on the X chromosome. It is twice as prevalent in males as in females. About one-third of individuals with the syndrome show features of autism, including trouble interacting with others, maintaining eye contact and learning to speak.
Studies have shown that mice lacking FMRP, or fragile X mental retardation protein, have immature dendritic spines, the neuronal branches that receive signals from other cells2,3, and too little of some synaptic proteins4.
The new study shows that FMRP regulates these synaptic proteins by blocking the action of ribosomes. Ribosomes are cellular structures that translate messenger RNA (mRNA), the transcript of genetic information, into protein. FMRP acts like a brake on this process, limiting the amount of protein produced by its targets.
As many of these targets help regulate brain development and plasticity, their misregulation may underlie symptoms of fragile X syndrome and autism, researchers say.
If FMRP is not available to repress translation at the right time and place, cells may make too much of its targets, says Jennifer Darnell, research associate professor of molecular neuro-oncology at Rockefeller University in New York City and co-investigator on the study. That excessive translation could lead to the defects of synaptic plasticity — the ability of synapses to regulate the strength of their connections — seen in both disorders.
“Loss of function of FMRP leads to so many defects of synaptic plasticity,” says Darnell. “The question is what subset of FMRP targets are responsible for the mouse and human phenotypes [of fragile X syndrome and autism]. It’s unlikely to be all 842.”
Hitchhiker protein:
The researchers used a technique called cross-linking immunoprecipitation (CLIP) to identify FMRP’s targets in the mouse brain. CLIP creates cross-links, or chemical bonds between proteins, using ultraviolet light.
The work is done in live mice 2 to 4 weeks of age, a period of great synaptic formation and synaptic plasticity.
The technique preserves physiological interactions in the cell, making the results more reliable, says Kimberly Huber, associate professor of neuroscience at the University of Texas Southwestern Medical Center, who was not involved with the research.
“Doing the ultraviolet cross-linking in vivo really increases the confidence that these are direct in vivo targets of FMRP,” she says. “Knowing what FMRP is binding to and potentially regulating is key to understanding how the gene is regulating brain function.”
Based on CLIP assays, other RNA-binding proteins seem to prevent translation only at specific locations on the mRNA, Darnell says. FMRP, by contrast, is a bit of a hitchhiker, hopping onto the ribosome in order to stall it, and interacting with mRNA at various points along its coding sequence.
“It has no specific binding site that we can find,” says Darnell. “That was a complete surprise.”
The CLIP assay identified 842 protein targets for FMRP. When the researchers compared these to a database of 117 genes associated with autism, they found a significant overlap of 28 genes. The list includes such well-studied candidate genes as SHANK3, PTEN, TSC2, and mGluR5.
Proteins at both the transmitting and receiving ends of the synapse turned up in the assay. This is surprising, says Darnell, as the loss of FMRP was thought to be most important at the post-synaptic end. “But FMRP is really regulating a very large percentage of both the pre- and post-synaptic proteome,” she says.
Huber says the study’s findings illuminate at least one long-standing puzzle in fragile X syndrome. Although researchers have long known that FMRP suppresses translation, the mechanism by which FMRP carried out this task was unknown.
The new study not only identifies a plethora of targets for FMRP, but also provides strong biochemical data about how FMRP stalls ribosomes and suppresses translation, she says. “Providing that rigorous data is a great contribution.”
References:
-
Darnell J.C. et al. Cell 146, 247-261 (2011) Abstract
-
Irwin S.A. et al. Am. J. Med. Genet. 98, 161-167 (2001) PubMed
-
Galvez R. and W.T. Greenough Am. J. Med. Genet. A. 135, 155-160 (2005) PubMed
-
Schütt J. et al. J. Biol. Chem. 284, 25479-25487 (2009) PubMed
Recommended reading
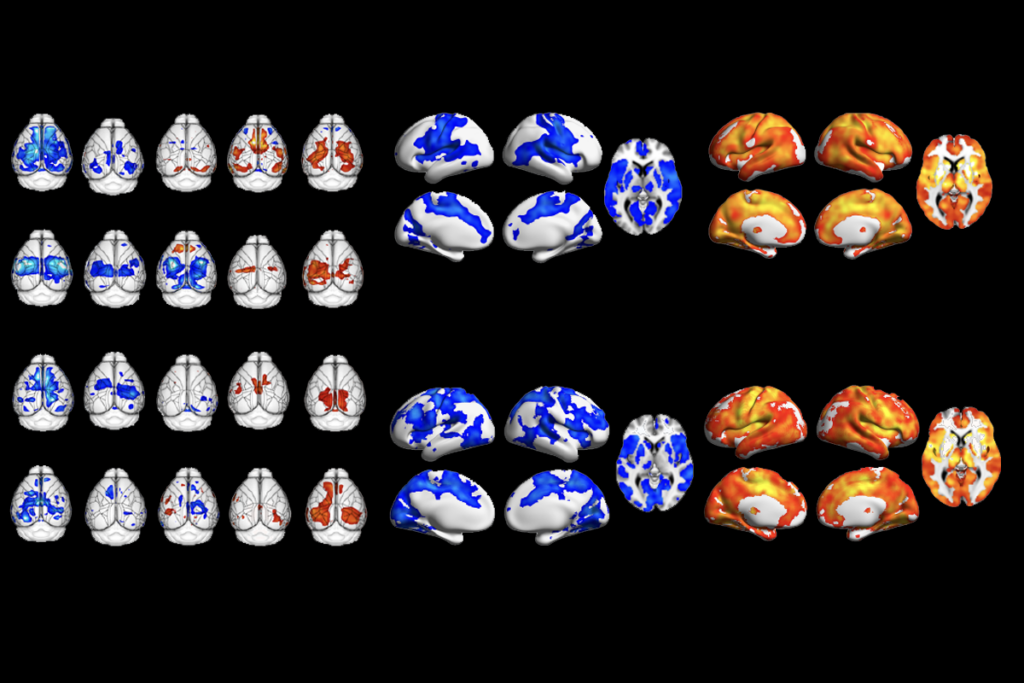

Too much or too little brain synchrony may underlie autism subtypes
Developmental delay patterns differ with diagnosis; and more
Split gene therapy delivers promise in mice modeling Dravet syndrome
Explore more from The Transmitter
During decision-making, brain shows multiple distinct subtypes of activity
Basic pain research ‘is not working’: Q&A with Steven Prescott and Stéphanie Ratté
