Researchers release first data from human connectome project
The first wave of data from the Human Connectome Project, a five-year $30 million effort to map the structure of the human brain, is now freely available, researchers announced at the 2012 Society for Neuroscience annual meeting in New Orleans.
The first wave of data from the Human Connectome Project, a five-year, $30 million effort to map the structure of the human brain, is now freely available, researchers announced at the 2012 Society for Neuroscience annual meeting in New Orleans.
In addition to imaging data, the researchers plan to release algorithms — such as the pulse sequences used in magnetic resonance imaging (MRI) — and other protocols to the public, which could be a boon to scientists studying autism. The researchers spent the first two years of the project perfecting imaging methods, and just began scanning the first participants in August.
For example, they have developed and evaluated new algorithms for cleaning up noisy imaging data, aiming to make the scans more sensitive.
Given the heterogeneity of autism, “you can’t just scan 20 people and capture the abnormalities,” says David Van Essen, one of the project’s lead investigators and professor of anatomy and neurobiology at Washington University in St. Louis. “You need methods that are sensitive enough to detect differences in individuals.”
Though the connectome project is limited to healthy adults, the tools and pipeline developed for the project could be applied to other groups, such as individuals with autism and children.
“We would love to have a developmental brain connectome,” says Story Landis, director of the National Institute of Neurological Disorders and Stroke, which funded part of the project. “But we just don’t have the funding.”
Big data:
When complete, the project will house structural and functional imaging data from 1,200 healthy adults, aged 22 to 35 years, along with information from behavioral tests, demographics and genetic testing. Each participant goes to Washington University for two days of brain scans and other tests.
The researchers plan to scan most of the participants using a 3 Tesla scanner, but a subset of participants will be scanned using a 7 Tesla machine, which has a stronger magnet and can therefore provide higher-resolution data.
They also plan to use magnetoencephalography, or MEG, which can track faster changes in brain activity than functional MRI can. (Functional MRI detects blood flow as an indirect measure of the brain’s electrical activity, whereas MEG measures electrical activity more directly.)
“There is a team of physicists across the world dedicated to optimizing [these approaches], and it has had a significant impact on the quality of the data,” says Heidi Johansen- Berg, a senior research fellow at Oxford University in the U.K. who is also part of the project. “We’re privileged as neuroscientists to have this.”
Researchers have so far scanned 36 people and released data on 12. “The purpose of this initial early data release is not that we expect to make major discoveries,” says Van Essen. “But because of the high quality of the data, we want the community to start working with these datasets.”
The researchers expect to release data on an additional 80 participants this winter. “By then, we hope to have the first generation of data-mining capabilities,” Van Essen says. They plan to release data quarterly, along with improvements to the data-mining platform.
Because the information will be made available to the public, the team has had to develop efficient ways to both store it and make it easily accessible.
“The data file per person is 30 gigabytes, so it’s not whipping across the Internet quickly,” says Van Essen.
The team has created a visualization platform, specialized for the multimodal nature of the project, that people can use to access the data.
Simultaneous slices:
One of the advances to come out of the project, developed by the University of Minnesota’s Kamil Ugurbil, is a protocol for scanning multiple slices of the brain at a time — typical methods do just one. That dramatically speeds up the process, which has both practical and scientific benefits.
A second arm of the project, at Massachusetts General Hospital and the University of California, Los Angeles, is developing new methods for diffusion tensor imaging, a variation of MRI that maps the brain’s wiring.
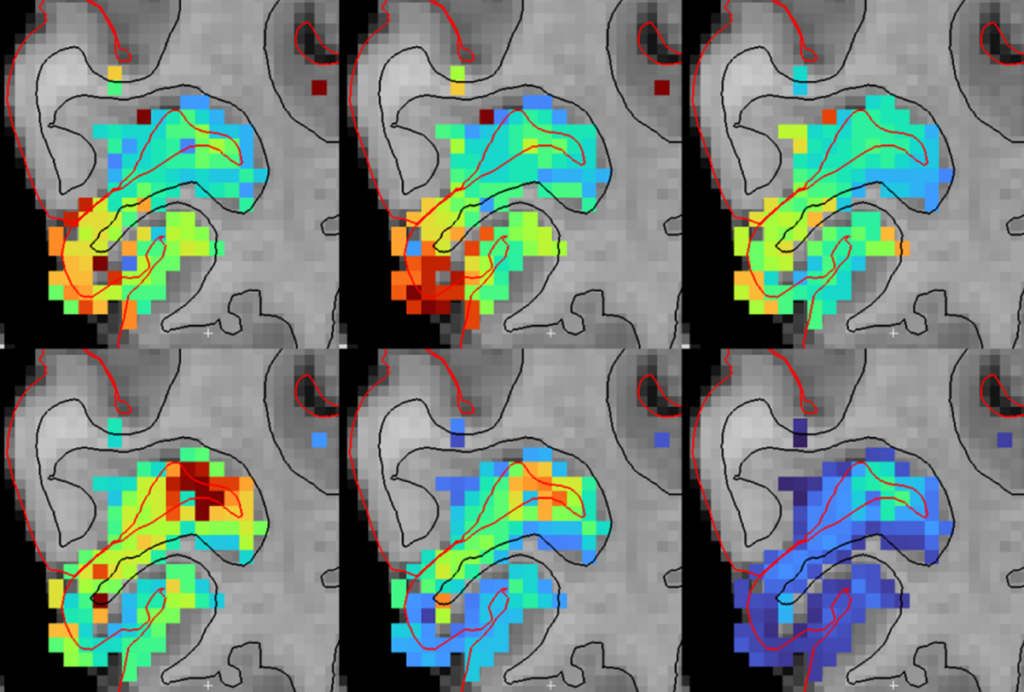
Each participant in the project undergoes both a structural and functional connectivity scan. In the latter, researchers predict functional connections by looking for correlations in activity among different brain regions. Because they collect and analyze the scans in a coordinated way, researchers can compare connectivity maps from the same person using both modalities.
“We see some overlap, but we also see some functional connections that we don’t see in the structure data,” says Van Essen. “It could be because functional imaging shows some connections that aren’t directly anatomical or it could be due to limitations in the imaging.”
Some of the tools that might be particularly useful for autism research are methods for correcting for movement in functional connectivity imaging. “This is a problem we’ve known about for years, but the magnitude of the problem has only recently been apparent,” says Van Essen.
If a participant moves more in the scanner, it appears as if they have more short-range connections. This is especially problematic in autism because a shortage of long-range connections and an abundance of short-range connections have been proposed to underlie some of the deficits associated with the disorder. Researchers disagree over whether findings from existing studies show a real difference or merely an artifact.
A number of methods have been developed for correcting the problem, and Van Essen says they are evaluating some of these as part of the project’s pipeline, which they also plan to release publicly.
For more reports from the 2012 Society for Neuroscience annual meeting, please click here.
Recommended reading
Developmental delay patterns differ with diagnosis; and more
Split gene therapy delivers promise in mice modeling Dravet syndrome
Changes in autism scores across childhood differ between girls and boys
Explore more from The Transmitter

‘Natural Neuroscience: Toward a Systems Neuroscience of Natural Behaviors,’ an excerpt