SFARI is pleased to announce that it expects to have funded 22 grants in response to the Explorer Awards request for applications (RFA) this year.

SFARI is pleased to announce that it expects to have funded 22 grants in response to the Explorer Awards request for applications (RFA) this year.

Frank McCormick, Ph.D., F.R.S., is the David A. Wood Distinguished Professor of Tumor Biology and Cancer Research at the University of California, San Francisco (UCSF) Helen Diller Family Comprehensive Cancer Center, and the leader of the RAS Initiative at the National Cancer Institute. His current research focuses on the understanding of RAS GTPases and how they can be therapeutically targeted in RAS-driven cancers, which are some of the most commonly occurring and most difficult to treat.

Paul Sternberg and colleagues establish an initial pipeline in C. elegans to screen autism-associated missense mutations for functional effects.

New Simons Searchlight data were recently added to SFARI Base. This data release included phenotypic data from individuals with 16p11.2 copy number variants (CNVs), 1q21.1 CNVs, 7q11.23 duplication and variants in 32 single genes associated with autism and related neurodevelopmental conditions.

SFARI is helping to make zebrafish models of high-risk autism genes available to the research community.
SFARI is now curating a set of zebrafish lines to study autism spectrum disorder. This includes mutant lines for 12 ASD risk genes, four of which are currently available to researchers and eight that will be available later this year.
A new collaboration between SFARI and the Nancy Lurie Marks Family Foundation will generate hundreds of induced pluripotent stem cells from individuals with autism and related neurodevelopmental conditions. These cell lines will become available to researchers starting next year.

New Simons Searchlight data were recently added to SFARI Base. This data release included phenotypic data from individuals with 16p11.2 copy number variants (CNVs), 1q21.1 CNVs, 7q11.23 duplication and variants in 29 single genes associated with autism and related neurodevelopmental conditions.
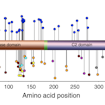
Kurt Haas and colleagues used multiple models and bioassays to assess the functional impact of more than 100 missense and nonsense mutations in PTEN, allowing for high-confidence predictions of pathogenicity.