
New phenotypic data from Simons Searchlight participants were recently added to SFARI Base. This release includes data from individuals with 94 gene changes and 15 copy number variants known to be connected to autism.

New phenotypic data from Simons Searchlight participants were recently added to SFARI Base. This release includes data from individuals with 94 gene changes and 15 copy number variants known to be connected to autism.

New Simons Searchlight data were recently added to SFARI Base. The data released included phenotypic data from individuals with 16p11.2 copy number variant (CNVs), 1q21.1 CNVs, 7q11.23 duplication and variants in 32 single genes associated with autism and related neurodevelopmental conditions.
New zebrafish lines with mutations in the high-confidence autism risk genes DYRK1A and GRIN2B have been added to SFARI resources.

New phenotypic data from Simons Searchlight participants were recently added to SFARI Base. This release includes data from individuals with 59 gene changes and nine copy number variants known to be connected to autism.
On March 25 and 28, 2022 SFARI hosted a workshop to explore the role of mitochondria in ASD risk. The workshop brought together experts in mitochondrial research spanning cellular, molecular, genetic, clinical and pharmacological areas and provided points for discussion on how to move this research forward.

New Simons Searchlight data were recently added to SFARI Base. This data release included phenotypic data from individuals with 16p11.2 copy number variants (CNVs), 1q21.1 CNVs, 7q11.23 duplication and variants in 32 single genes associated with autism and related neurodevelopmental conditions.
Zebrafish lines for autism risk genes DYNC1H1 and MEF2C have been recently added to SFARI resources to study autism spectrum disorder.

New Simons Searchlight data were recently added to SFARI Base. This data release included phenotypic data from individuals with 16p11.2 copy number variants (CNVs), 1q21.1 CNVs, 7q11.23 duplication and variants in 32 single genes associated with autism and related neurodevelopmental conditions.
SFARI is pleased to announce that it intends to fund 19 grants in response to the Summer 2020 Pilot Award request for applications.
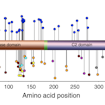
Kurt Haas and colleagues used multiple models and bioassays to assess the functional impact of more than 100 missense and nonsense mutations in PTEN, allowing for high-confidence predictions of pathogenicity.